Main menu
Common skin conditions
NEWS
Join DermNet PRO
Read more
Quick links
Authors: Dr Jenna E. Rayner, Dermatology Registrar; Dr David L. Duffy, Statistical Geneticist; Dr Richard A. Sturm, Principal Research Fellow, Dermatology Research Centre, University of Queensland Diamantina Institute, University of Queensland, Brisbane, Queensland, Australia. DermNet Editor in Chief: Adjunct A/Prof Amanda Oakley, Dermatologist, Hamilton, New Zealand. Copy edited by Gus Mitchell. January 2020.
Introduction - genes Introduction - IRF4 gene Variants IRF4-related phenotypes Clinical features Diagnosis Patient management
The physical features of hair, skin, and eye colour, freckling, melanocytic naevi and the skin’s response to the sun are influenced by major gene polymorphisms.
Sun exposure stimulates tanning, where melanocytes produce the biopolymer melanin and transfer it to keratinocytes.
The interferon regulatory factor 4 (IRF4) gene belongs to a family of DNA-binding proteins that play an important role in immune system development and function [1].
IRF4 polymorphisms have been independently associated with pigmentation and age-specific effects on melanocytic naevus count in European-derived populations [2,3].
A single nucleotide polymorphism (SNP) within intron 4 of IRF4, rs12203592*C/T, plays a major role in pigmentation.
IRF4 is one of only 2 genes, along with tyrosinase (TYR), known to affect the skin, eye, hair, and together with the melanocortin 1 receptor (MC1R), freckling. IRF4 independently affects total body naevus counts [4].
IRF4 is predominantly expressed in blood cells with key roles in lymphoid, myeloid and dendritic cell lineages and adipocytes. IRF4 is also expressed in melanocytes in naevi and primary melanomas [5,6].
Asian and African populations are fixed for the ancestral rs12203592*C allele, being monomorphic. In European populations, the rs12203592*T allele is found at around 17% allele frequency. This allele frequency is 40% in Ireland.
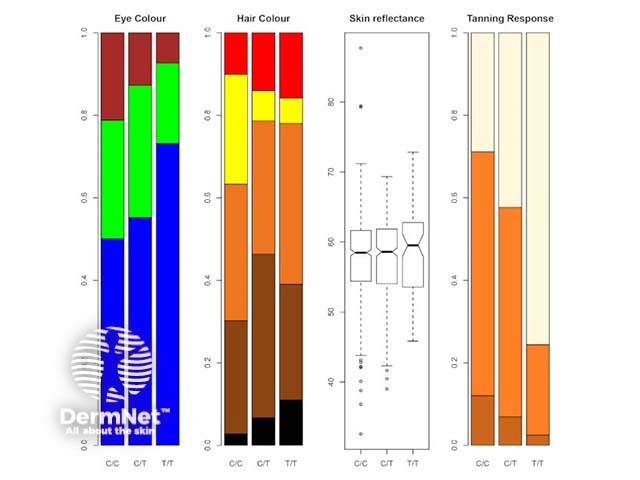
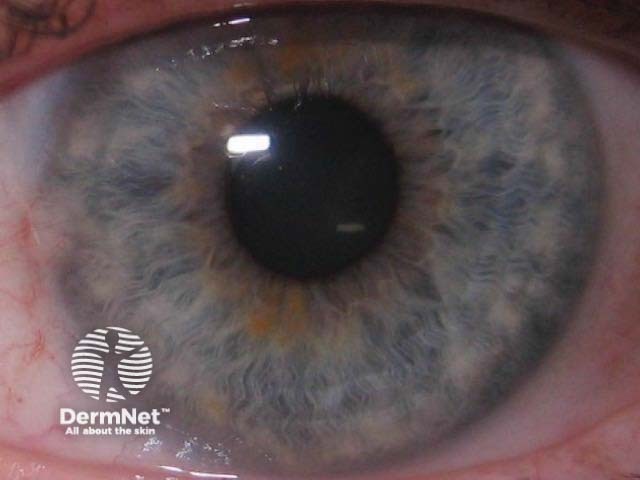
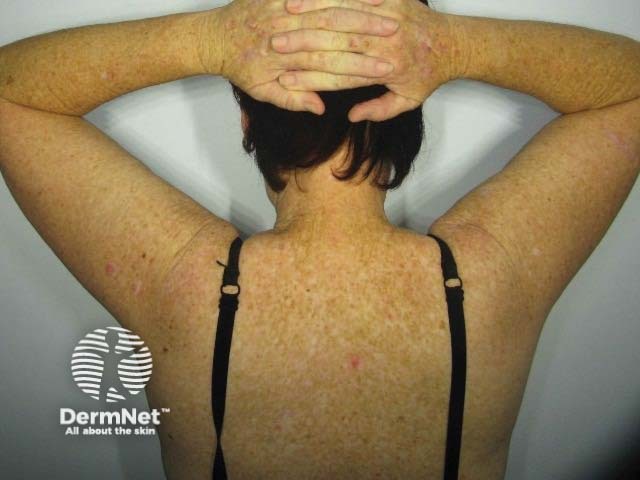
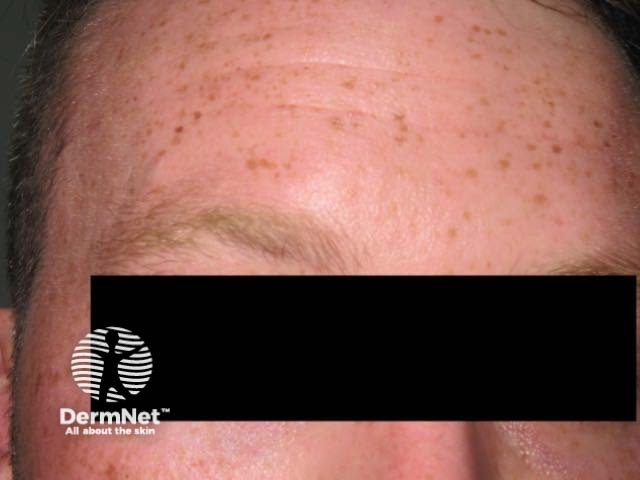
The rs12203592*T allele was statistically and significantly associated with dark hair colour, light eye colour, and decreased ability to tan [2,3,7]. Figure 1 shows the frequency of this phenotype by rs12203592 genotype in 1200 individuals from South-east Queensland. Figure 2, 3 and 4 demonstrate the blue eye and freckling phenotype of IRF4 rs12203592*T/T patients. IRF4 has also been one of the few genes linked to hair greying [8].
A strong genotype-by-age interaction for total body naevus count has also been reported for this SNP, which is also a predictor of melanoma risk (Table 1). Compared to rs12203592*C/C or C/T carriers, homozygous T/T individuals have higher naevus counts as adolescents, which reverses with age so that adults have lower naevus counts. The rs12203592*T allele was also associated with decreased counts of raised (papular and dermal) melanocytic naevi and increased counts of flat (compound and junctional) naevi [3].
The Study of Nevi in Children (SONIC) identified five significant associations (p < 0.005) between IRF4 rs12203592 variants and the clinical appearance of naevi. SNPs in IRF4 showed significant associations with globular patterns [9]. The IRF4 rs12203952 polymorphism was very strongly associated with naevus counts (p < 0.0001). T alleles, compared with C, are linked to increased naevus counts at any level of sun exposure.
IRF4 rs12203592*C= 83% |
IRF4 rs12203592*T =17% |
Low adolescent naevus count a, e, f |
High adolescent naevus count a, e, f |
High adult naevus count a, e, f |
Low adult naevus count a, e, f |
Darker skin colour b |
Lighter skin colour b |
Brown/dark eye colour b |
Blue/light eye colour b, g |
Better tanning b |
Reduced tanning b |
Melanoma naevus remnant b |
Melanoma solar elastosis b |
Melanoma with lower Breslow thickness c |
Melanoma with greater Breslow thickness c |
Increased survival in melanoma d |
Reduced survival in melanoma d |
Lower incidence of nodular melanoma h |
Higher incidence of nodular melanoma h |
Table 1: rs12203592*T carriers have a reduced number of naevi with age, a fairer phenotype and an increased melanoma Breslow thickness with localised chronic UV solar damage.
a (Duffy et al. 2010) [3], b (Gibbs et al. 2016) [7], c (Gibbs et al. 2017) [10], d (Potrony et al. 2017) [11], e (Kvaskoff et al. 2011) [12], f (Orlow et al. 2015) [9], g (Pena-Chilet et al. 2013) [13], h (Rayer et al. 2019) [16]
Genetic testing for IRF4 variants is most often undertaken for research purposes and forensic analysis of DNA samples [14]. As of 2019, the entire coding region is assessed on an Illumina Infinium HumanCoreExome-24 Microarray or through a bi-directional Sanger sequence analysis of intron 4.
People likely to carry IRF4 rs12203592*T variants have a phenotype which demonstrates high adolescent naevus count, lighter skin and eye colour, and a reduced ability to tan. These patients’ melanomas are frequently associated with solar elastosis, increased Breslow thickness (Figure 3), and reduced survival in melanoma.
Sun protection includes wearing sun-protective clothing when outdoors, applying sunscreen at regular intervals, and avoiding intense or frequent sunburn.
People carrying the IRF4 variant gene should be advised to arrange skin checks by a trained health professional. Self skin examination education should emphasise lesions that are enlarging or changing in colour, shape and structure. Awareness of the ABCDE features of melanoma can be useful; these features are:
The proposed addition of “EFG” (Elevated, Firm and Growing for more than a month) to the ABCD acronym aids clinical assessment of lesions. This may be particularly relevant for patients with an IRF4 rs12203592*T/T genotype, whose melanomas are associated with an increased Breslow thickness [15].